An enzyme that digests plastic could boost recycling – A MILLION plastic bottles are sold every minute. Many are not recycled and of those that are, only a small fraction become bottles again – Enzyme digests plastic recycling - Arhive
Enzyme digests plastic recycling Enzyme digests plastic recycling Enzyme digests plastic recycling Enzyme digests plastic recycling Enzyme digests plastic recycling Enzyme digests plastic recycling Enzyme digests plastic recycling Enzyme digests plastic recycling Enzyme digests plastic recycling Enzyme digests plastic recycling Enzyme digests plastic recycling
An enzyme that digests plastic could boost recycling
Auf Wiedersehen, PET
 A MILLION plastic bottles are sold every minute. Many are not recycled and of those that are, only a small fraction become bottles again. That is, in part, because recycling polyethylene terephthalate (PET), the polymer used to make such bottles, back into material robust enough to hold, say, a fizzy drink, is hard. What would be helpful is a way to break down PET into the chemicals that made it in the first place. These could then be used to make new high-grade PET.
A MILLION plastic bottles are sold every minute. Many are not recycled and of those that are, only a small fraction become bottles again. That is, in part, because recycling polyethylene terephthalate (PET), the polymer used to make such bottles, back into material robust enough to hold, say, a fizzy drink, is hard. What would be helpful is a way to break down PET into the chemicals that made it in the first place. These could then be used to make new high-grade PET.
This week John McGeehan of the University of Portsmouth, in Britain, and his colleagues report details of a bacterial enzyme called “PETase” that can do just that. Furthermore, they have engineered a version of this enzyme that can digest plastic faster than the natural variety. Their work is published in the Proceedings of the National Academy of Sciences.
Latest stories
-
An enzyme that digests plastic could boost recycling
-
James Comey against the president
-
The business of death is changing around the world
-
A plan to put beds on planes
-
Why some countries still ban gay men from giving blood
-
Fixing the flaws in today’s capitalism
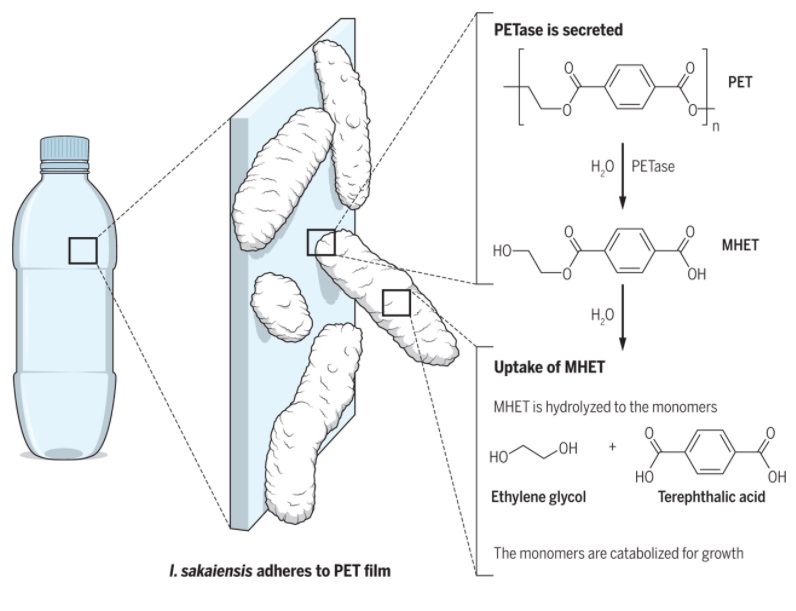
PETase is secreted by a plastic-munching bacterium called Ideonella sakaiensis 201-F6. This bug was discovered in 2016 at a PET-bottle recycling plant in Sakai, Japan. The researchers behind its discovery showed that the enzyme degrades PET into mono(2-hydroxyethyl) terephthalic acid (MHET). A second enzyme then breaks MHET down further, into terephthalic acid and ethylene glycol. The bacterium then uses these chemicals as food sources. The discoverers of PETase also suggested that it may have evolved from bacterial enzymes used to break down cutin, a waxy polymer that coats leaves. That is, in itself, remarkable—for PET has been used widely only since the 1970s, meaning that the enzyme must have evolved to do its job within the past 50 years.
I. sakaiensis digests PET far too slowly, however, to be of much use for industrial recycling of the plastic. To make it so requires understanding how the enzymes do their work. This is what Dr McGeehan and his colleagues set out to do. As MHET is far easier to break down by standard chemical means than PET, they focused on PETase.
They compared the DNA sequence of the PETase gene to that of cutinases from thousands of species of bacteria, looking for systematic differences. They then created new versions of PETase, each with one or more of its amino-acid building blocks changed to resemble those of ancestral cutinases.
As many of the differences between PETase and cutinases were, presumably, what allowed the PETase to do its job, they expected these new enzymes to digest the plastic less efficiently. To their surprise, however, one of the engineered enzymes (with two amino acids mutated to be more cutinase-like) was able to digest PET about 20% faster than the natural one. That is a modest increase, but one that came about by accident rather than design. This, Dr McGeehan argues, shows there is plenty of scope for further improvement.
The team determined the structures of their enzymes by protein crystallography, a technique that takes detailed pictures of a molecule by bombarding crystals of it with X-rays (in this case, at the Diamond Light Source, a machine in Oxfordshire that produces particularly strong X-rays for such purposes). They then used computer modelling to look at how a molecule of PET might dock with the enzyme’s active site—the region where the chemical reaction that breaks down the plastic actually occurs. The more-efficient enzyme they engineered appears to hold the plastic molecule more snugly in the active site than the naturally occurring version.
Interesting though all this is, there is still much to do before PETase can become a useful enzyme. At the moment, a litre of a solution of even the improved enzyme would break down just a few milligrams of plastic per day. Its plastic-digesting ability must therefore be improved by a hundredfold or more to be commercially useful.
This the team hopes to do, in part, by using clues from the enzyme’s structure. Further improvements could come by designing the enzyme to work at temperatures above 70ºC, when PET becomes rubbery, and thus more easily digestible. Bacteria that live in hot springs, and that have cutinases that function at such temperatures, might be pressed into service here. The gene for the enzyme would also have to be transplanted into bacteria that can be grown easily at industrial scales. If these hurdles can be surmounted, though, PETase might make a dent in the scourge of plastic waste.
